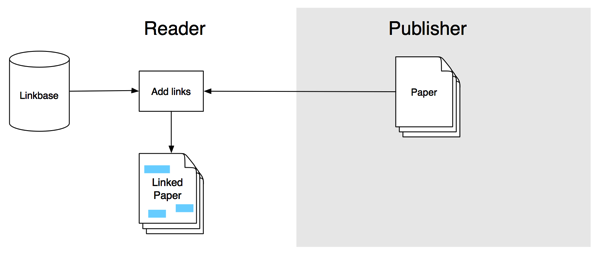
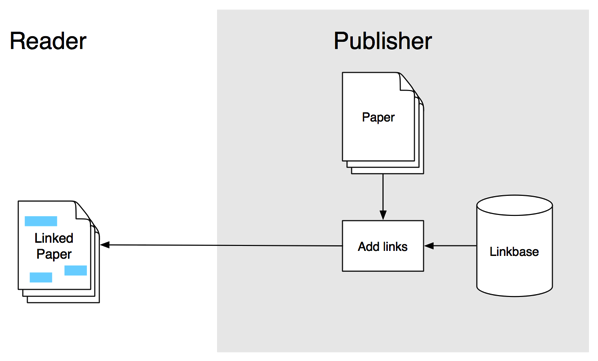
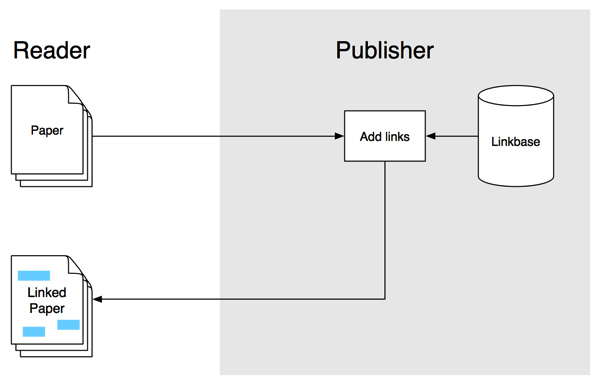
In the previous post I sketched out a workflow to annotate articles using hypothes.is and aggregate those annotations. I threw this together for the hackday at ReCon 16 in Edinburgh, and the hack day gave me a chance to (a) put together a crude visualisation of the aggregated annotations, and (b) recruit CrossRef's Rachael Lammey (@rachaellammey) into doing some annotations as well so I could test how easy it was to follow my garbled instructions and contribute to the project.
We annotated the paper A new species of shrew (Soricomorpha: Crocidura) from West Java, Indonesia (doi:10.1644/13-MAMM-A-215). If you have the hypothes.is extension installed you will see our annotations on that page, if not, you can see them using the hypothes.is proxy: https://via.hypothes.is/http://dx.doi.org/10.1644/13-MAMM-A-215.
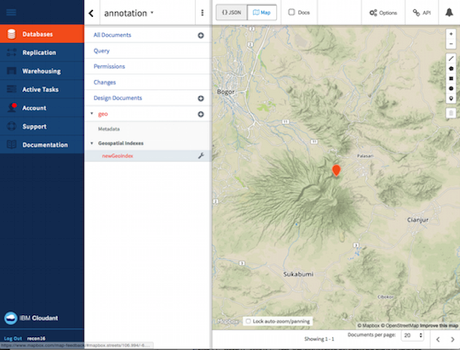
Rachael and I both used the IFTTT tool to send our annotations to a central store. I then created a very crude summary page for those annotations: http://bionames.org/~rpage/recon16-annotation/www/index.html?id=10.1644/13-MAMM-A-215. When this page loads it queries the central store for annotations on the paper with DOI 10.1644/13-MAMM-A-215, then creates some simple summaries.
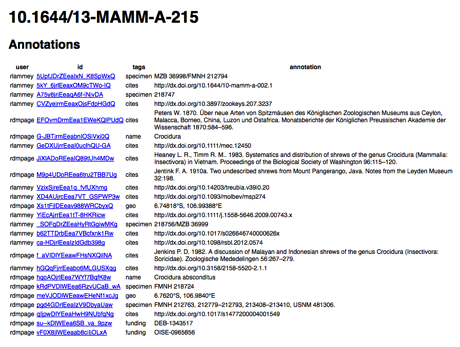
For example, here is a list of the annotations. The listed is "typed" by tags, that is, you can tell the central store what kind of annotation is being made using the "tag" feature in hypothes.is. On this example, we've picked out taxonomic names, citations, geographical coordinates, specimen codes, grants, etc.

Given that we have latitude and longitude pairs, we can generate a map:
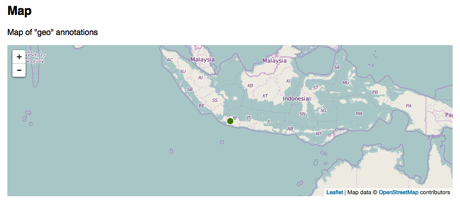
The names of taxa can be enhanced by adding pictures, so we have a sense of what organisms the paper is about:

The metadata on the web page for this article is quite rich, and hypothes.is does a nice job of extracting it, so that we have a list of DOIs for many of the articles this paper cites. I've chosen to add annotations for articles that lack DOIs but which may be online elsewhere (e.g., BioStor).

What's next
This demo shows that it's quite straightforward to annotate an article and pull those annotations together to create a central database that can generate new insights about a paper. For example, we can generate a map even if the original paper doesn't provide one. Conversely, we could use the annotations to link entities such as museum specimens to the literature that discusses those specimens. Given a specimen code in a paper we could look up that code in GBIF (using GBIF's API, or a tool like "Material Examined", see Linking specimen codes to GBIF). Hence we could go from code in paper to GBIF, or potentially from GBIF to the paper that cites the specimen. Having a central annotation store potentially becomes a way to build a knowledge graph linking different entities that we care about.Of course, a couple of people manually annotating a few papers isn't scalable, but because hypothes.is has an API we can scale this approach (for another experiment see Hypothes.is revisited: annotating articles in BioStor). For example, we have automated tools to locate taxonomic names in text. Imagine that we use those tools to create annotations across the accessible biodiversity literature. We can then aggregate those into a central store and we have an index to the literature based on taxonomic name, but we can also click on any annotation and see that name in context as an annotation on a page. We could manually augment those annotations, if needed, for example by correcting OCR errors.
I think there's scope here for unifying the goals of indexing, annotation, and knowledge graph building with a fairly small set of tools.
 I will be at
I will be at